Mendelian Randomization (MR) Analysis: Principles, Advantages, Limitations and Practical R Implementation
Mendelian Randomization (MR) analysis is a powerful epidemiological method that uses genetic variants (single nucleotide polymorphisms, SNPs) as instrumental variables (IVs) to infer the causal relationship between an exposure factor and an outcome. It overcomes the limitations of traditional observational studies in dealing with confounding factors and reverse causality, and has been widely used in the study of causal associations between complex traits (e.g., allergic diseases) and diseases (e.g., cancer), as demonstrated in the research on allergic rhinitis (AR) and cancer risk. This article will elaborate on the core principles, advantages, practical limitations of MR analysis, and provide a detailed, reproducible R code implementation based on a two-sample MR study of AR and cancers.
1. Core Principles of MR Analysis
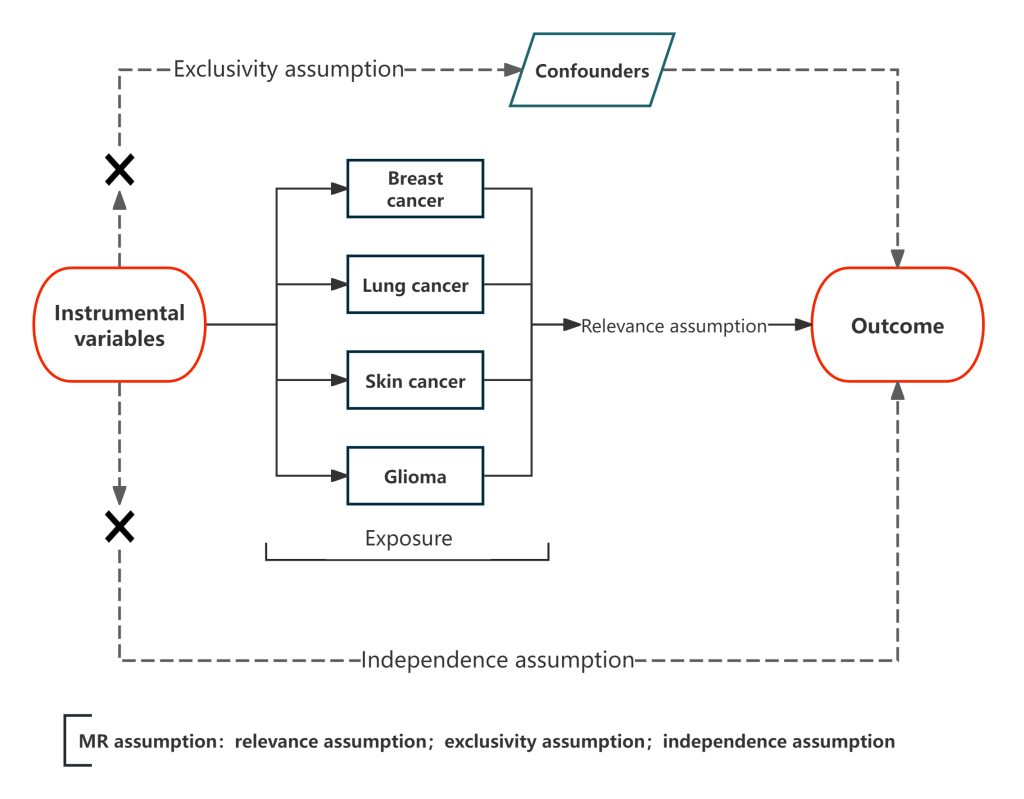
- Relevance assumption: IVs (genetic variants) are significantly associated with the exposure factor (e.g., SNPs are closely linked to the genetic susceptibility of AR).
- Independence assumption: IVs are independent of any confounding factors that affect both the exposure and the outcome (e.g., SNPs for AR are not associated with smoking, age, etc., which affect cancer risk).
- Exclusivity assumption: IVs affect the outcome only through the exposure factor, and there is no other independent causal pathway (e.g., AR-related SNPs affect cancer risk only by regulating AR, not by other biological mechanisms).
Two-sample MR: The Most Commonly Used MR Design
- The exposure GWAS dataset: Contains genetic variation and exposure phenotype data (e.g., AR GWAS data from 112,583 European participants).
- The outcome GWAS dataset: Contains genetic variation and outcome phenotype data (e.g., breast cancer, lung cancer GWAS data from different European cohorts).
2. Key Advantages of MR Analysis
2.1 Avoid confounding bias fundamentally
2.2 Resolve the problem of reverse causality
2.3 Make full use of public GWAS resources
2.4 Support multi-method verification and improve result reliability
3. Limitations of MR Analysis (Methodological + Practical Application)
3.1 Inherent methodological limitations
(1) Risk of weak instrumental variables
(2) Horizontal pleiotropy
(3) Linkage disequilibrium (LD) interference
(4) Limitations of statistical methods
3.2 Practical application limitations (Reflected in the AR-cancer study)
(1) Population stratification and poor generalizability
(2) Uneven statistical power of different outcomes
- Sufficient power: ER- breast cancer (80.00%), skin cancer (SC, 77.20%);
- Moderate power: Lung squamous cell carcinoma (LUSC, 65.90%), basal cell carcinoma (BCC, 58.00%);
- Insufficient power: Lung adenocarcinoma (LUAD, 9.60%), ER+ breast cancer (43.90%), glioma (44.10%).
(3) Lack of stratified analysis and prospective verification
(4) Inability to analyze gene-environment interaction
4. Standard Analysis Workflow of Two-sample MR
- Data acquisition: Obtain GWAS summary data of exposure and outcome from public databases (IEU OpenGWAS, GWAS Catalog, FinnGen, etc.), and ensure the consistency of population ancestry.
- IVs screening: Strictly screen SNPs according to the criteria (genome-wide significance P<5e−8, clumping to remove LD, Steiger Test to verify causal direction, exclude SNPs associated with outcomes/confounders, RadialMR to remove abnormal IVs).
- IVs strength evaluation: Calculate the F-statistic of the screened SNPs to ensure no weak IVs (F>10).
- Core MR analysis: Use the main method (IVW) for primary analysis, and multiple robust methods (MR-Egger, Weighted median, Weighted mode) and novel methods (cML-MA, ConMix, MR-RAPS, dIVW) for cross-verification.
- Sensitivity analysis:
- Cochran Q test: Evaluate the heterogeneity of IVs;
- MR-Egger-intercept + MR-PRESSO: Detect horizontal pleiotropy;
- Leave-one-out analysis: Verify whether the results are driven by a single abnormal SNP.
- Result visualization: Draw forest plots, scatter plots, leave-one-out plots to intuitively display the causal effect and sensitivity analysis results.
- Result interpretation: Combine statistical power, population characteristics, and biological mechanisms to interpret the results, and clarify the limitations of the study.
5. Practical R Code Implementation of Two-sample MR
TwoSampleMR (for MR data processing, analysis and visualization) and MRPRESSO (for horizontal pleiotropy detection). The following code is a complete, reproducible two-sample MR analysis code based on the study, taking allergic rhinitis (AR) as exposure and ER- breast cancer as outcome as an example, and the code can be directly modified for other exposure-outcome combinations.5.1 Environment Construction
# Set working directory
setwd("your_working_directory")
# Install packages (run only once)
install.packages(c("TwoSampleMR", "MRPRESSO", "tidyverse", "ggplot2"))
devtools::install_github("MRCIEU/TwoSampleMR") # Install the latest version of TwoSampleMR
# Load packages
library(TwoSampleMR)
library(MRPRESSO)
library(tidyverse)
library(ggplot2)
# Set seed for reproducibility
set.seed(123)
5.2 Data Acquisition
- AR:
ukb-b-7178(IEU OpenGWAS) - ER- breast cancer:
ieu-a-1166(IEU OpenGWAS)
# Obtain exposure data (AR)
exposure_dat <- extract_instruments(
id = "ukb-b-7178", # AR dataset ID
p1 = 5e-8, # Genome-wide significance threshold
clump = TRUE, # Open clumping to remove LD
clump_kb = 10000, # Clumping kb threshold
clump_r2 = 0.001, # Clumping r2 threshold
clump_p = 1,
pop = "EUR" # European ancestry
)
# Obtain outcome data (ER- breast cancer)
outcome_dat <- extract_outcome_data(
snps = exposure_dat$SNP, # SNPs from exposure data
id = "ieu-a-1166", # ER- breast cancer dataset ID
pop = "EUR"
)
# Harmonize exposure and outcome data (unify allele direction, remove incompatible SNPs)
mr_dat <- harmonise_data(
exposure_dat = exposure_dat,
outcome_dat = outcome_dat,
action = 2 # Remove SNPs with allele direction inconsistency
)
5.3 IVs Strength Evaluation (Calculate F-statistic)
Calculate the R2 and F-statistic of each SNP to verify that there are no weak IVs (F>10). The formula is from the AR-cancer study:
R2=β2×2×EAF×(1−EAF)
F=R2×(N−K−1)÷[K×(1−R2)]
- β: Regression coefficient of SNP-exposure association;
- EAF: Effect allele frequency;
- N: Sample size of exposure GWAS;
- K: Number of IVs (SNPs).
# Add EAF and sample size (AR sample size N=112583)
mr_dat$eaf <- mr_dat$exposure_eaf
mr_dat$N <- 112583
K <- nrow(mr_dat)
# Calculate R2 and F-statistic
mr_dat <- mr_dat %>%
mutate(
R2 = (exposure_beta^2) * 2 * eaf * (1 - eaf),
F = R2 * (N - K - 1) / (K * (1 - R2))
)
# View IVs strength (F-statistic >10 is strong IVs)
print(paste("Number of final IVs:", K))
print(round(mr_dat[, c("SNP", "F")], 2))
print(paste("Minimum F-statistic:", min(mr_dat$F)))
5.4 Core MR Analysis
# Core MR analysis (traditional methods)
mr_results <- mr(mr_dat, method_list = c(
"ivw_fixed", # Fixed effects IVW (main method)
"mr_egger_regression", # MR-Egger
"weighted_median", # Weighted median
"weighted_mode" # Weighted mode
))
# Convert beta to OR (OR = exp(beta), 95% CI = exp(beta ± 1.96*se))
mr_results_or <- mr_results %>%
mutate(
OR = exp(b),
OR_lower = exp(b - 1.96 * se),
OR_upper = exp(b + 1.96 * se),
p_value = pval
) %>%
select(method, OR, OR_lower, OR_upper, p_value)
# View results
print(round(mr_results_or, 4))
# Debiased IVW (dIVW, novel method) - eliminate weak IVs bias
divw_results <- mr_dwive(mr_dat)
divw_or <- data.frame(
method = "debiased_ivw",
OR = exp(divw_results$b),
OR_lower = exp(divw_results$b - 1.96 * divw_results$se),
OR_upper = exp(divw_results$b + 1.96 * divw_results$se),
p_value = divw_results$pval
)
print(round(divw_or, 4))
5.5 Sensitivity Analysis
#### 5.5.1 Heterogeneity test (Cochran Q)
hetero_results <- mr_heterogeneity(mr_dat)
print(hetero_results[, c("method", "Q", "Q_pval")])
#### 5.5.2 Horizontal pleiotropy test
# MR-Egger-intercept
pleio_egger <- mr_pleiotropy_test(mr_dat)
print(paste("MR-Egger-intercept P-value:", round(pleio_egger$pval, 4)))
# MR-PRESSO (detect and correct abnormal IVs)
presso_dat <- mr_dat %>%
select(SNP, beta.exposure = exposure_beta, se.exposure = exposure_se,
beta.outcome = outcome_beta, se.outcome = outcome_se)
presso_results <- MRPRESSO(presso_dat, NbDistribution = 1000, SignifThreshold = 0.05)
print(presso_results$MainResults) # Global test P-value
print(presso_results$OutlierTest) # Abnormal SNP detection
#### 5.5.3 Leave-one-out analysis
leave1out_results <- mr_leaveoneout(mr_dat)
# Visualize leave-one-out results
p_leave1out <- mr_leaveoneout_plot(leave1out_results)
print(p_leave1out)
ggsave("leaveoneout_plot.pdf", p_leave1out, width = 10, height = 6)
5.6 Result Visualization
#### 5.6.1 Forest plot (OR and 95% CI of causal effect)
p_forest <- mr_forest_plot(mr_results, mr_dat)
print(p_forest)
ggsave("mr_forest_plot.pdf", p_forest, width = 12, height = 8)
#### 5.6.2 Scatter plot (SNP level exposure-outcome association)
p_scatter <- mr_scatter_plot(mr_results, mr_dat)
print(p_scatter[[1]])
ggsave("mr_scatter_plot.pdf", p_scatter[[1]], width = 8, height = 6)
#### 5.6.3 Funnel plot (detect publication bias)
p_funnel <- mr_funnel_plot(mr_singlesnp(mr_dat))
print(p_funnel)
ggsave("mr_funnel_plot.pdf", p_funnel, width = 8, height = 6)
5.7 Extended Analysis (Novel Methods)
- cML-MA: Implemented through the
MRcMLpackage (GitHub: https://github.com/QingyuanYin/MRcML) - ConMix: Implemented through the
MRConMixpackage (GitHub: https://github.com/mrcieu/MRConMix) - MR-RAPS: Implemented through the
MRrapspackage (CRAN)
6. Summary and Research Suggestions
MR analysis is a revolutionary epidemiological method for causal inference, which has become an important tool for studying the causal association between complex traits and diseases by using genetic variants as instrumental variables. The two-sample MR study of AR and cancers fully demonstrates the application value of MR analysis: it clarifies that genetically predicted AR is a risk factor for ER- breast cancer and a protective factor for LUSC, SC and BCC, and provides a reliable causal basis for subsequent biological mechanism research.
- Strictly screen IVs: Follow the three basic assumptions, use clumping to remove LD, Steiger Test to verify causal direction, and F-statistic to exclude weak IVs.
- Multi-method cross-verification: Combine traditional methods (IVW, MR-Egger) and novel methods (cML-MA, MR-RAPS) for analysis, and take the results with consistent directions of multiple methods as the final conclusion.
- Comprehensive sensitivity analysis: Use Cochran Q, MR-Egger-intercept, MR-PRESSO and leave-one-out analysis to evaluate heterogeneity and horizontal pleiotropy, and remove abnormal IVs.
- Pay attention to statistical power: For outcomes with low statistical power, expand the sample size or combine multiple GWAS datasets for meta-analysis to avoid false negative results.
- Strengthen the combination with other studies: MR analysis provides genetic causal evidence, and the combination with prospective cohort studies, in vitro cell experiments and in vivo animal experiments can further verify the causal relationship and explore the underlying biological mechanisms.
- Expand the research population: Collect GWAS data of different ethnic groups to improve the generalizability of the research results.

